Interactive jobs
From HP-SEE Wiki
CMSLTM
To run interactive jobs, one needs to run the "nrniv" executable with the "-mpi" parameter. Interactive jobs are only used to test that the simulation can run without problems with the given number of processes. For production runs, jobs should be added to the PBS queue.
Our version of NEURON, compiled using gcc with openMPI support is installed in /home/gkastel/src/nrn-7.1/x86_64/. The mpiexec launcher must be used to launch the neuron executable on multiple nodes in the cluster. This constitutes an interactive job.
Non-interactive jobs are the same as interactive ones, except that they are launched by the PBS system and not by the user, and their output is redirected to output files in the launch directory instead of standard output.
Running interactive jobs did not require any modifications in the application.
Example run of interactive jobs:
mpiexec -np 10 nrniv -dll "$NLIBS" -mpi finalpar.hoc
Deep sequencing for short fragment alignment (DeepAligner)
Section contributed by OU & SZTAKI
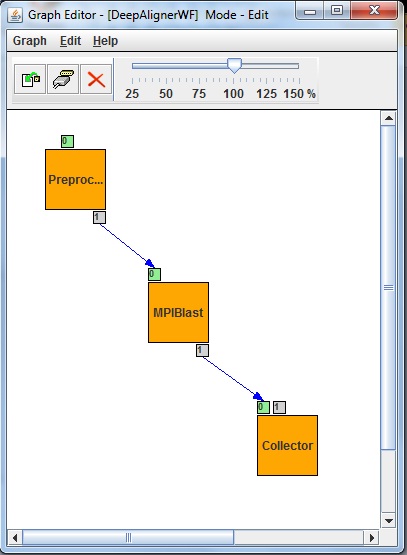
The DeepAligner application’s workflow has been developed as a Parameter Study workflow with usage of autogenerator port (second small box around left top box in Fig xs1.) and collector job (right bottom box in Fig. xs1). The preprocessor job generates a set of input files from some pre-adjusted parameter. Then the second job (middle box in Fig. x1) will be executed as many times as the input files specify. The second job is an MPI based BLAST executable (MPIBlast) which aligns short sequences. The inputs of the MPI job are the sets of sequences (defined by the researcher) and the already deployed sequence database fragments. The last job of the workflow is a Collector which is used to collect several files and then process them as a single input. Collectors force delayed job execution until the last file of the input file set to be collected has arrived to the Collector job. The workflow engine computes the expected number of input files at run time. When all the expected inputs arrived to the Collector it starts to process all the incoming inputs files as a single input set. Finally output files will be generated, and will be stored on a Storage Element of the DCI shown as little box around the Collector in Fig 1.
Fig 1: DeepAligner workflow with the MPI based Blast job in the middle More information: [1]
In-silico Disease Gene Mapper
Section contributed by OU & SZTAKI
The in-silico Disease Gene Mapper (DiseaseGeneMapper) was ported successfully with the workflow based gUSE development environment. The ported DiseaseGeneMapper application is operated as an on-line service on the HP-SEE’s Bioinformatics eScience Gateway.
DiseaseGene (In-silico Disease Gene Mapper) [2]