MDSCS
From HP-SEE Wiki
(→Running on Several HP-SEE Centres) |
(→Publications) |
||
| (39 intermediate revisions not shown) | |||
| Line 5: | Line 5: | ||
* Scientific contact: ''Dr. Armen Poghosyan, sicnas@sci.am'' | * Scientific contact: ''Dr. Armen Poghosyan, sicnas@sci.am'' | ||
* Technical contact: ''Dr. Hrachya Astsatryan, hrach@sci.am'' | * Technical contact: ''Dr. Hrachya Astsatryan, hrach@sci.am'' | ||
| - | * Developers: ''National Academy of Sciences of the Republic of Armenia'' | + | * Developers: ''[http://www.sci.am/ National Academy of Sciences of the Republic of Armenia]'' |
* Web site: [http://grid.am/index.php/Special:AGApplications?action=7 Bioinformatics] Group | * Web site: [http://grid.am/index.php/Special:AGApplications?action=7 Bioinformatics] Group | ||
== Short Description == | == Short Description == | ||
| - | The parallel molecular dynamics simulation of complex system consisting of surfactant/polymer mixtures is planning to carry out. The self –assembling of surfactant systems is also planning to study. The molecular dynamics | + | The parallel molecular dynamics simulation of complex system consisting of surfactant/polymer mixtures is planning to carry out. The self –assembling of surfactant systems is also planning to study. The molecular dynamics results together with experimental finding will help to understand the mechanism of interactions in physical point of view and, which is most important, will give us an important information concerning the conformation and localization of system components (polymer, surfactant, water, ect.). The main research method is the molecular dynamics simulation, as well as for comparison the GPS measurement and X-ray methods are planning to use. The application based on [http://www.ks.uiuc.edu/Research/namd/ NAMD] and [http://www.gromacs.org GROMACS] packages, which are designed for high-performance simulation of large molecular systems. |
== Problems Solved == | == Problems Solved == | ||
| - | The 50ns | + | The 50ns molecular dynamics simulation run of SDS/water/oil system in presence of polyampholyte at pH=9 have been done. It is planned to continue the additional 50ns simulation run of mentioned system at pH=4. The analysis of molecular dynamics data is done for the system at pH=9. The self-organization of surfactant/water system is in progress (currently at ~16ns run point) |
== Scientific and Social Impact == | == Scientific and Social Impact == | ||
| Line 22: | Line 22: | ||
== Collaborations == | == Collaborations == | ||
| - | * Institut für Chemie of Universität Potsdam, Germany | + | * [http://www.chem.uni-potsdam.de/ Institut für Chemie of Universität Potsdam], Potsdam, Germany |
| - | * International Center for Theoretical Physics, Italy | + | * [http://www.ictp.it/ International Center for Theoretical Physics], Trieste, Italy |
== Beneficiaries == | == Beneficiaries == | ||
| - | * | + | * Main beneficiaries are research groups investigating complex systems of molecular dynamics consisting of surfactant/polymer mixtures and micelles |
== Number of users == | == Number of users == | ||
| Line 34: | Line 34: | ||
== Development Plan == | == Development Plan == | ||
| - | * Concept: '' | + | * Concept: ''Done before the project started'' |
| - | * Start of alpha stage: '' | + | * Start of alpha stage: ''Done before the project started'' |
| - | * Start of beta stage: '' | + | * Start of beta stage: ''M1-M3'' |
| - | * Start of testing stage: '' | + | * Start of testing stage: ''M3-M6'' |
| - | * Start of deployment stage: '' | + | * Start of deployment stage: ''M6-M10'' |
| - | * Start of production stage: '' | + | * Start of production stage: ''M11-M24'' |
== Resource Requirements == | == Resource Requirements == | ||
| - | * Number of cores required for a single run: '' | + | * Number of cores required for a single run: ''64-2048'' |
| - | * Minimum RAM/core required: '' | + | * Minimum RAM/core required: ''4GB'' |
* Storage space during a single run: ''0.5TB'' | * Storage space during a single run: ''0.5TB'' | ||
* Long-term data storage: ''5TB'' | * Long-term data storage: ''5TB'' | ||
| - | * Total core hours required: '' | + | * Total core hours required: ''more than 2 000 000'' |
== Technical Features and HP-SEE Implementation == | == Technical Features and HP-SEE Implementation == | ||
| Line 53: | Line 53: | ||
* Primary programming language: ''C/C++ '' | * Primary programming language: ''C/C++ '' | ||
* Parallel programming paradigm: ''MPI/OpenMP '' | * Parallel programming paradigm: ''MPI/OpenMP '' | ||
| - | * Main parallel code: ''NAMD,GROMACS'' | + | * Main parallel code: ''NAMD, GROMACS'' |
* Pre/post processing code: ''VMD package'' | * Pre/post processing code: ''VMD package'' | ||
| - | * Application tools and libraries: '' | + | * Application tools and libraries: ''NAMD, GROMACS, FFTW'' |
== Usage Example == | == Usage Example == | ||
| Line 63: | Line 63: | ||
== Infrastructure Usage == | == Infrastructure Usage == | ||
| - | * Home system: '' | + | * Home system: ''ArmGrid'' |
** Applied for access on: ''07.2011'' | ** Applied for access on: ''07.2011'' | ||
** Access granted on: ''07.2011'' | ** Access granted on: ''07.2011'' | ||
| - | ** Achieved scalability: '' | + | ** Achieved scalability: ''256 cores'' |
| - | * Accessed production systems: | + | * Accessed production systems: ''BG/BG'' |
| - | + | ** Applied for access on: ''07.2011'' | |
| - | + | ** Access granted on: ''07.2011'' | |
| - | + | ** Achieved scalability: ''2048 cores'' | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
== Running on Several HP-SEE Centres == | == Running on Several HP-SEE Centres == | ||
| - | * Benchmarking activities and results: ''The benchmarking of mentioned system was done using 512,1024,2048 and 4096 processors on BlueGene/P'' | + | * Benchmarking activities and results: ''The benchmarking of mentioned system was done using 512,1024,2048 and 4096 processors on BlueGene/P, Bulgaria.'' |
== Achieved Results == | == Achieved Results == | ||
| + | * Molecular Dynamics simulation run of SDS/water/oil system in presence of polyampholyte at pH=9 | ||
| + | [[File:MDSCS.jpg]] | ||
| + | |||
| + | * Sodium dodecyl sulfate (SDS)/water system at 45ns checkpoint | ||
| + | [[File:milions_at45ns.JPG]] | ||
== Publications == | == Publications == | ||
| + | * Armen H. Poghosyan, Levon H. Arsenyan, Aram A. Shahinyan, [http://pubs.acs.org/doi/abs/10.1021/la302378r?prevSearch=Molecular%2BDynamics%2BStudy%2Bof%2BIntermediate%2BPhase%2Bof%2BLong%2BChain%2BAlkyl%2BSulfonate%252FWater%2BSystems&searchHistoryKey= Molecular Dynamics Study of Intermediate Phase of Long Chain Alkyl Sulfonate/Water Systems], Langmuir Journal, American Chemical Society, DOI: 10.1021/la302378r | ||
* A. Poghosyan, L. Arsenyan, H. Astsatryan, M. Gyurjyan, H. Keropyan and A. Shahinyan, [http://www.scirp.org/journal/PaperDownload.aspx?paperID=17494&returnUrl=http%3a%2f%2fwww.scirp.org%2fjournal%2fPaperInformation.aspx%3fpaperID%3d17494/NAMD Package Benchmarking on the Base of Armenian Grid Infrastructure], Journal of Communications and Network, Scientific Research Publishing, Vol. 4 No. 1, 2012, pp. 34-40, doi:10.4236/cn.2012.41005 | * A. Poghosyan, L. Arsenyan, H. Astsatryan, M. Gyurjyan, H. Keropyan and A. Shahinyan, [http://www.scirp.org/journal/PaperDownload.aspx?paperID=17494&returnUrl=http%3a%2f%2fwww.scirp.org%2fjournal%2fPaperInformation.aspx%3fpaperID%3d17494/NAMD Package Benchmarking on the Base of Armenian Grid Infrastructure], Journal of Communications and Network, Scientific Research Publishing, Vol. 4 No. 1, 2012, pp. 34-40, doi:10.4236/cn.2012.41005 | ||
| - | * A.H. Poghosyan, L. H. Arsenyan, H.V. Astsatryan, [http://www.mipro.hr/MIPRO2012/ELink.aspx/Comparative NAMD Benchmarking of Complex System on Bulgarian BlueGene/P], IEEE Proceedings of MIPRO 2012 - Jubilee 35th International Convention, Opatija, Croatia | + | * A. Shahinyan, P. Hakobyan, L. Arsenyan, A. Poghosyan, The study of lyotropic liquid crystal structure using the molecular dynamics method, [http://www.tandfonline.com/toc/gmcl20/current/ Journal of Molecular Crystals and Liquid Crystals], 2012, DOI:10.1080/15421406.2012.687174 |
| + | * A.H. Poghosyan, L. H. Arsenyan, H.V. Astsatryan, [http://www.mipro.hr/MIPRO2012/ELink.aspx/Comparative NAMD Benchmarking of Complex System on Bulgarian BlueGene/P], IEEE Proceedings of MIPRO 2012 - Jubilee 35th International Convention, Opatija, Croatia, pp. 319-321. | ||
| + | * A.A.Shahinyan, L.H.Arsenyan, A.H.Poghosyan, [http://lrb.jinr.ru/conf/mssmbs12/BookOfAbstracts-MSSMBS2012.pdf The Study of Phase Diagram of the Surfactant/Water System by Molecular Dynamics Simulation], Book of Abstracts of 5th Japan – Russia International Workshop "Molecular Simulation Studies in Material and Biological Sciences", Joint Institute for Nuclear Research, Dubna, Russia, September 9 – 12, 2012. | ||
* A.H.Poghosyan, L.H. Arsenyan, A.A. Shahinyan "Molecular dynamics study of long chain alkyl sulfonate/water system", [http://www.journals.elsevier.com/journal-of-colloid-and-interface-science/ JCIS] ''in review'' | * A.H.Poghosyan, L.H. Arsenyan, A.A. Shahinyan "Molecular dynamics study of long chain alkyl sulfonate/water system", [http://www.journals.elsevier.com/journal-of-colloid-and-interface-science/ JCIS] ''in review'' | ||
== Foreseen Activities == | == Foreseen Activities == | ||
| + | |||
| + | * Micelle self-assemble | ||
Latest revision as of 17:59, 6 January 2013
General Information
- Application's name: Molecular Dynamics Study of Complex Systems (MDSCS)
- Virtual Research Community: Life Sciences
- Scientific contact: Dr. Armen Poghosyan, sicnas@sci.am
- Technical contact: Dr. Hrachya Astsatryan, hrach@sci.am
- Developers: National Academy of Sciences of the Republic of Armenia
- Web site: Bioinformatics Group
Short Description
The parallel molecular dynamics simulation of complex system consisting of surfactant/polymer mixtures is planning to carry out. The self –assembling of surfactant systems is also planning to study. The molecular dynamics results together with experimental finding will help to understand the mechanism of interactions in physical point of view and, which is most important, will give us an important information concerning the conformation and localization of system components (polymer, surfactant, water, ect.). The main research method is the molecular dynamics simulation, as well as for comparison the GPS measurement and X-ray methods are planning to use. The application based on NAMD and GROMACS packages, which are designed for high-performance simulation of large molecular systems.
Problems Solved
The 50ns molecular dynamics simulation run of SDS/water/oil system in presence of polyampholyte at pH=9 have been done. It is planned to continue the additional 50ns simulation run of mentioned system at pH=4. The analysis of molecular dynamics data is done for the system at pH=9. The self-organization of surfactant/water system is in progress (currently at ~16ns run point)
Scientific and Social Impact
The investigations will help to understand the mechanism of interaction of anionic SDS and cationic/anionic/noncharged polymer and to receive important information on the dynamical and structural features of mentioned systems. The results of the proposed investigations will make an important contribution to basic researches in Colloid Physics.
Collaborations
- Institut für Chemie of Universität Potsdam, Potsdam, Germany
- International Center for Theoretical Physics, Trieste, Italy
Beneficiaries
- Main beneficiaries are research groups investigating complex systems of molecular dynamics consisting of surfactant/polymer mixtures and micelles
Number of users
20
Development Plan
- Concept: Done before the project started
- Start of alpha stage: Done before the project started
- Start of beta stage: M1-M3
- Start of testing stage: M3-M6
- Start of deployment stage: M6-M10
- Start of production stage: M11-M24
Resource Requirements
- Number of cores required for a single run: 64-2048
- Minimum RAM/core required: 4GB
- Storage space during a single run: 0.5TB
- Long-term data storage: 5TB
- Total core hours required: more than 2 000 000
Technical Features and HP-SEE Implementation
- Primary programming language: C/C++
- Parallel programming paradigm: MPI/OpenMP
- Main parallel code: NAMD, GROMACS
- Pre/post processing code: VMD package
- Application tools and libraries: NAMD, GROMACS, FFTW
Usage Example
job_script using Grid (JDL), Grid Site (PBS) or BlueGeneP
Infrastructure Usage
- Home system: ArmGrid
- Applied for access on: 07.2011
- Access granted on: 07.2011
- Achieved scalability: 256 cores
- Accessed production systems: BG/BG
- Applied for access on: 07.2011
- Access granted on: 07.2011
- Achieved scalability: 2048 cores
Running on Several HP-SEE Centres
- Benchmarking activities and results: The benchmarking of mentioned system was done using 512,1024,2048 and 4096 processors on BlueGene/P, Bulgaria.
Achieved Results
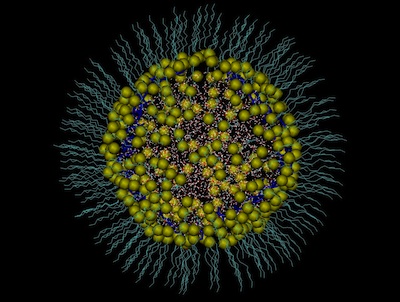
- Molecular Dynamics simulation run of SDS/water/oil system in presence of polyampholyte at pH=9
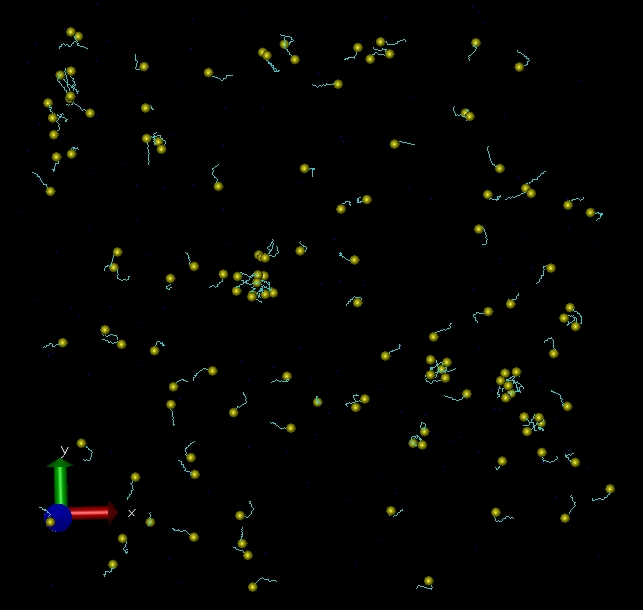
- Sodium dodecyl sulfate (SDS)/water system at 45ns checkpoint
Publications
- Armen H. Poghosyan, Levon H. Arsenyan, Aram A. Shahinyan, Molecular Dynamics Study of Intermediate Phase of Long Chain Alkyl Sulfonate/Water Systems, Langmuir Journal, American Chemical Society, DOI: 10.1021/la302378r
- A. Poghosyan, L. Arsenyan, H. Astsatryan, M. Gyurjyan, H. Keropyan and A. Shahinyan, Package Benchmarking on the Base of Armenian Grid Infrastructure, Journal of Communications and Network, Scientific Research Publishing, Vol. 4 No. 1, 2012, pp. 34-40, doi:10.4236/cn.2012.41005
- A. Shahinyan, P. Hakobyan, L. Arsenyan, A. Poghosyan, The study of lyotropic liquid crystal structure using the molecular dynamics method, Journal of Molecular Crystals and Liquid Crystals, 2012, DOI:10.1080/15421406.2012.687174
- A.H. Poghosyan, L. H. Arsenyan, H.V. Astsatryan, NAMD Benchmarking of Complex System on Bulgarian BlueGene/P, IEEE Proceedings of MIPRO 2012 - Jubilee 35th International Convention, Opatija, Croatia, pp. 319-321.
- A.A.Shahinyan, L.H.Arsenyan, A.H.Poghosyan, The Study of Phase Diagram of the Surfactant/Water System by Molecular Dynamics Simulation, Book of Abstracts of 5th Japan – Russia International Workshop "Molecular Simulation Studies in Material and Biological Sciences", Joint Institute for Nuclear Research, Dubna, Russia, September 9 – 12, 2012.
- A.H.Poghosyan, L.H. Arsenyan, A.A. Shahinyan "Molecular dynamics study of long chain alkyl sulfonate/water system", JCIS in review
Foreseen Activities
- Micelle self-assemble