HP-SEE Wiki
From HP-SEE Wiki
Virtual Research Communities
Computational Chemistry VRC
Overview
Computational chemistry and material science is one of the highlighted research areas in computational science and a typical heavy user of HPC resources. The computational technologies are an indispensable tool for investigations in domains like quantum molecular dynamics, molecular modelling, nano-technology and design of new materials. Considering the size of the problems to be studied, the required calculations are extremely computationally intensive. Thus HPC would greatly facilitate the proposed work allowing the researchers to deal not only with “pilot” or model systems but to work on big and complicated real systems, which are physically and technologically more significant and challenging. These studies will extend understanding of some fundamental science issues and are of practical importance for pharmaceutical industry, nanotechnology, biomedicine, and many others.
Initially Computational Chemistry VRC supports 7 applications with main developers in 6 SEE countries, collaborating with scientists from more than 20 advanced research centers in Europe.
Computational Chemistry Applications
| Acronym | Application Name | Main developer | Stage |
| CFDOF | CFD Analysis of Combustion | Faculty of Mech. Engineering, University of Banja Luka (UoBL), Bosnia - Herzegovina | testing 09.2011 |
| CompChem | Quantum Mechanical, Molecular Mechanics, and Molecular Dynamics computation in chemistry | Univeristy of Belgrade, Faculty of Chemistry, Republic of Serbia | production 09.2011 |
| FMD-PA | Design of fullerene and metal-diothiolene-based materials for photonic applications | Computational Chemistry Group of NHRF, Greece | production 08.2011 |
| HC-MD-QM-CS | Hybrid Classical/Quantum Molecular Dynamics – Quantum Mechanical Computer Simulation of Condensed Phases | UKIM, Institute of Chemistry, Faculty of Natural Science and Mathematics, FYROM | production 08.2011 |
| ISyMAB | Integrated System for Modeling and data Analysis of complex Biomolecules | IFIN-HH/DPETI, Romania | production 05.2012 |
| MDCisplatin | Molecular Design of Platinum Group Metal Complexes as Potential Non-classical Cisplatin Analogues | Acad. Roumen Tsanev Institute of Molecular Biology-BAS, Bulgaria | production 07.2012 |
| PCACIC | Principal component analysis of the conformational interconversions in large-ring cyclodextrins | IOCCP-BAS, Bulgaria | production 08.2011 |
Computational Physics VRC
Overview
Computational Physics is nowadays the main beneficiary of the scientific HPC, large-scale numerical computations being necessary whenever the complexity of the physical systems investigated does not allow the derivation of an analytical solution. The main objective of the Computational Physics VRC is to join together the various physics research teams from the SEE area and to provide them access to a powerful HPC infrastructure and tools which will make possible their participation in multidisciplinary and international collaborations. For this purpose, software developers from 7 countries (Albania, Bosnia and Herzegovina, Bulgaria, FYR of Macedonia, Republic of Moldova, Romania, Serbia) contributed with 12 applications in the fields of High Energy and Particle Physics, Plasma Physics, Physics of Condensed Matter, Atomic Physics, and Computational Fluid Dynamics. The application range extends from nanoelectronics, micro-devices optimization and the modeling of robotic devices for biomedicine, to improved means for feature detection in satellite images, which leads to better mapping, localization and search services.
Computational Physics Applications
| Acronym | Application Name | Main developer | Stage |
| AMR_PAR | Parallel algorithm and program for the solving of continuum mechanics equations using Adaptive Mesh Refinement | Institute of Mathematics and Computer Science, Laboratory of Mathematical Modelling, Republic of Moldova | production 09.2011 |
| EagleEye | Feature Extraction from Satellite Images Using a Hybrid Computing Architecture | University Politehnica of Bucharest / Computer Science and Engineering, Romania | production 08.2011 |
| FAMAD | Fractal Algorithms for MAss Distribution | Institute for Space Sciences, Romania | testing 07.2011 |
| FuzzyCmeans | Parallel Fuzzy C Means for classification/Feature detection category | West University of Timisoara/Computer Science Department, Romania | production 08.2011 |
| GENETATOMICS | Genetic algorithms in atomic collisions | Institute of Physics, Faculty of Natural Science and Mathematics, UKIM, FYROM | production 09.2011 |
| GIM | Geophysical Inversion Modeling | Polytechnic University of Tirana, Albania | testing 05.2011 |
| HAG | High energy physics Algorithms on GPU | Institute for Space Sciences, Romania | testing 09.2011 |
| HMLQCD | Hadron Masses from Lattice QCD | University of Tirana, Albania | production 07.2011 |
| NUQG | Numerical study of ultra-cold quantum gases | Scientific Computing Laboratory, Institute of Physics Belgrade, Serbia | production 08.2011 |
| SET | Simulation of electron transport | Institute of Information and Communication Technologies, Bulgarian Academy of Sciences, Bulgaria | production 08.2011 |
| SFHG | Self Avoiding Hamiltonian Walk on Gaskets | Faculty of Mechanical Engineering, Dept. of Thermomechanics, University of Banja Luka, Bosnia and Herzegovina | 2012 |
| SIMPLE-TS_2D | Finite Volume Method for calculation of 2D gas-microflows using standard MPI | Kiril Stoyanov Shterev and Stefan Kanchev Stefanov, Institute of Mechanics – BAS / Department of “Complex and multiphase Flows”, Bulgaria | production 08.2011 |
Life Sciences VRC
Overview
Life Sciences depend heavily on the use of HPC for both data mining and data integration as well as for the simulation of biological systems. HPC technologies are essential for research areas such genome analysis, expression profiling, -omics analysis and biological simulations, whereby a vast amount of experimental data needs to be analyzed and synthesized into reasonable hypothesis. Thus HPC would greatly facilitate the various applications described in this project, enabling the respective research teams to study questions that have thus far been intractable due to their high computational complexity. The use of HPC in the Life Sciences applications with help further our understanding of basic problems in the fields of DNA sequence analysis, comparative genomics, and brain modeling among others and can be of great importance for the health sector.
The Life Sciences VRC supports 7 applications with main developers in 5 SEE countries (Greece, Hungary, Montenegro, Armenia, Georgia) working in the areas of computational biology, computational biophysics, DNA sequence analysis and computational genomics. The various projects involve collaborations with numerous scientists both in Europe and the U.S. and will foster the development of new collaborations among the participant SEE countries.
Life Sciences Applications
| Acronym | Application Name | Main developer | Stage |
| CMSLTM | Computational Models of Short and Long Term Memory | George Kastellakis, IMBB-FORTH, Greece | production 09.2011 |
| DeepAligner | Deep sequencing for short fragment alignment | Miklos Kozlovszky, Gergely Windisch, Judit Márton,Obuda University (OU), John von Neumann Faculty of Informatics, Biotech Group, Hungary | production 02.2012 |
| DiseaseGene | In-silico Disease Gene Mapper | Miklos Kozlovszky, Gergely Windisch, Judit Márton, Obuda University (OU), John von Neumann Faculty of Informatics, Biotech Group, Hungary | production 02.2012 |
| MC-WND | High resolution medical tissue image classifier | Miklos Kozlovszky, Gergely Windisch, Obuda University (OU), John von Neumann Faculty of Informatics, Biotech Group, Hungary | production 03.2013 |
| DNAMA | DNA Multicore Analysis | School of Computer & Communication Sciences, Laboratory for Computational Biology and Bioinformatics, RAxML software, Montenegro | production 11.2011 |
| MDSCS | Molecular Dynamics Study of Complex systems | Dr. Armen Poghosyan, Dr. Hrachya Astsatryan, National Academy of Sciences of the Republic of Armenia | production 07.2011 |
| miRs | Searching for novel miRNA genes and their targets | Anastasis Oulas, IMBB/FORTH, Greece | production 05.2011 |
| MSBP | Modeling of some biochemical processes with the purpose of realization of their thin and purposeful synthesis | Jumber Kereselidze, Tbilisi State University Department of Natural Science, Georgia | production 11.2011 |
Fast Track access to HP-SEE infrastructure for new user communities
Since 2012, a continuous call for proposals is opened for fast track access to HP-SEE resources, targeted at new user communities. Researchers from the South-East Europe region covered by the project can run their scientific applications on resources provided by HPC centers in Bulgaria, Serbia, and Romania, and benefit from the technical support of HP-SEE experts during a two- or three-months period. The resources offered are limited to 4 nodes (48-64 CPU cores) and 4x480 GPUs cores per run. Developers of Chemistry and Life Science applications are especially encouraged to apply.
Fast Track Applications
| Acronym | Application Name | Domain | Main developer | Stage |
| LNAWENR | Asteroseismic pipeline and Web based portal based on LNAWENR software code to be used in the KASC/KEPLER Space Mission | Space Science | Marian Doru SURAN, Astronomical Institute of the Romanian Academy, Romania | ended Dec. 2012 |
| SyMBA_GPU | Study of the initial formation stages of the solar system | Space science | Kleomenis TSIGANIS, Aristotle University of Thessaloniki, Greece | running |
| ASEP2D | Simulation of 2D Asymmetric Exclusion Processes | Computational Physics | Margarita IFTI, University of Tirana, Albania | ended June 2013 |
| SLVP | Selective Load Value Prediction | Engineering | Adrian FLOREA, 'Lucian Blaga' University of Sibiu, Romania | ended Aug. 2013 |
| NAMD | MelanoMD | Computational Chemistry | Adina MILAC, Institute of Biochemistry of the Romanian Academy, Romania | ended July 2013 |
| SeqSize | Sequencing Size and Power | Life Sciences | Dean PALEJEV, Bulgarian Academy of Sciences, Institute of Mathematics and Informatics, Bulgaria | ended Aug. 2013 |
| TMDC | Electronic structure, optical and transport properties of layered transition-metal dichalcogenides and their intercalated compounds | Computational Physics | Yurii CHOMAKOW, Institute of Applied Physics, Academy of Sciences of Moldova | ended July 2013 |
| PSoBBB | Population Status of Brown Bears in Bulgaria | Life Sciences | Anna JORDANOVA, Institute of Molecular Biology, Bulgarian Academy of Sciences | running |
| MUSICO | MUscle SImulation COde | Bioengineering | Djordje NEDIC, Faculty of Science, University of Kragujevac, Serbia | running |
User Guidelines
Getting Access to HP-SEE Infrastructure
Porting Support
Supporting Tools
Developer Guidelines
HPC programming techniques
Code programming and optimization
Architecture-specific optimization
- Porting between 32 and 64 bit addressing
- Adapting code to memory architecture
- Porting between different processor architectures
- Driver-level programming
Setting up the software environment
Choice and usage of libraries and application tools
Compiler-level optimization
Using development tools
Input/Output methodologies
Usage of job submission systems
Scalability testing and optimization
Benchmarking
HPC Infrastructure Operations
HPC Centers
Bulgaria
Hungary
- Resource centre Debrecen SC
- Resource centre NIIFI SC
- Resource centre Pecs SC
- Resource centre Szeged SC
Serbia
Romania
MKD
Core Services
Monitoring Tools
Network Operations
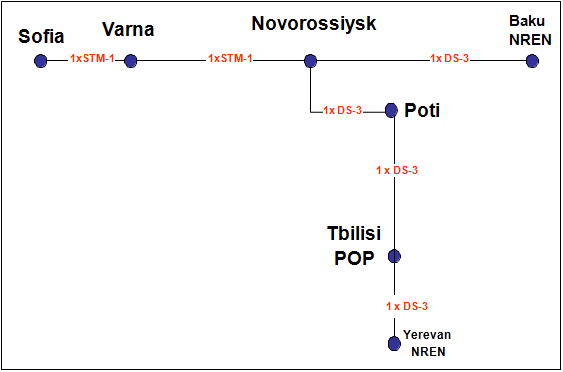
HP-SEE project connects South Caucasus National Research and Education Networks (NRENs), Azerbaijan Research & Educational Networking Association (AzRENA) and National Academy of Sciences of Armenia (NAS RA) to GEANT with 45 Mbps connections. These two NRENs were connected to GEANT at Q4 of 2011 and these connections will be valid during the project life time.
HP-SEE project built a virtual NOC composed of the NOC personnel from AzRENA, NAS RA and ULAKBIM and this virtual team operates the network. Network operations includes monitoring and management of Sofia-Baku and Sofia-Yerevan links.
NRENS
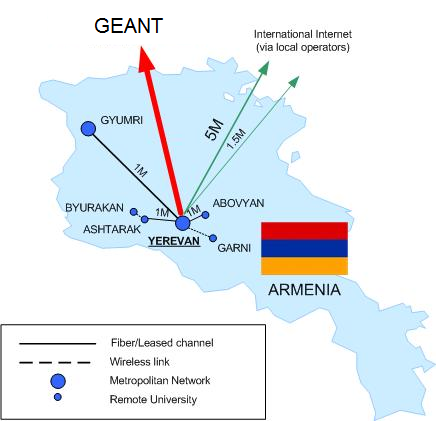
NAS RA
The backbone map of NAS RA is given in the following figure. This map basically shows the users of the GEANT link provided by HP-SEE project.
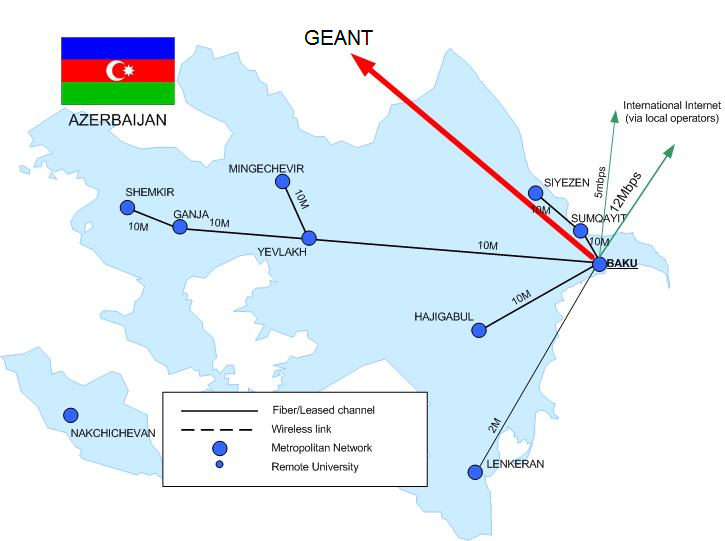
AzRENA
The backbone map of AzRENA is given in the following figure. This map basically shows the users of the GEANT link provided by HP-SEE project.
Monitoring and Management Tools
HP-SEE NOC uses Nagios, MRTG & RRDTool, CACTI and NfSEN for management and monitoring purposes and all of these tools are open-source. The findings of these tools and the usage graphics of HP-SEE link are available at: http://network.hp-see.eu
Nagios
Nagios is an open source network monitoring application which watches the hosts and services specified. Simple alerting mechanisms such as e-mail and sms are the equipments of Nagios to inform the NOC personnel in case of possible problems on the services or hosts being watched. In this deployment a PING service is associated with these defined hosts so that Nagios can continuously pole the link between ULAKBIM and the two NRENs.
MRTG and RRDTool
The Multi Router Traffic Grapher - MRTG is one of the most widely used to generate graphics of link usages. MRTG uses Simple Network Management Protocol (SNMP) to monitor network devices and draws pictures showing how much traffic has passed through each interface. Although MRTG generally uses SNMP as an input, it can also use other input methods such as output of files, plugins or custom scripts. HP-SEE monitor is equipped with MRTG and with the related configurations, it generates the graphics of Sofia-Baku and Sofia-Yerevan links in daily, weekly, monthly and yearly manner. In each 5 minutes, the usage statistics are gathered from AzRENA and NAS RA routers and the related graphics are updated.
WeatherMap
WeatherMap is a network visualization tool, uses data already collected from network devices and creates your network map. HP-SEE NOC uses WeatherMap to read the link usage data MRTG and generate HP-SEE links maps with link utilization ratios.
CACTI
CACTI is PHP based frontend by combining all the useful functionalities of RRDTool, SNMP and MRTG. HP-SEE NOC uses CACTI to aggregated data from Yerevan and Baku routers and to generate on demand graphs. HP-SEE NOC currently generates CPU usage, ping latency and traffic usage graphics via CACTI. All of these graphics are also generated in hourly, daily, weekly and monthly manner.
NfSen
NetFlow is a network protocol developed by Cisco Systems for collecting IP traffic information. NetFlow has become an industry standard for traffic monitoring. Today it is supported by almost all router manufactures providing similar functionality with a differed label and standards like sflow, ipfix. Open source flow generator software implementations such as softflowd, frobe can be used to generate a flow on Linux, FreeBSD, NetBSD and OpenBSD operating systems. In order to capture and store the generated flow in a analyzable format many flow capture and analyzer software were developed by open source community. NfSen, which was developed by GEANT2 project, is a graphical web based front end of nfdump and it is used to store and analyze network traffic flow traces in combination with NfDump. A flow trace is accounting information aggregated from the packets travelling over a router. Accounting information for all packets with the same 5-tuples (source IP address, source port number, destination IP address, destination port number and protocol) is aggregated under the same flow trace.
The HP-SEE Advanced Traffic Management System is installed by using two NfSen instances. One is used to capture the flows exported by Yerevan router and one is used to capture flow by AzRENA router. This system is a living one, which will be enhanced by addition of currently available NfSen plug-ins such as security related once in parallel to the growing needs of HP-SEE NOC.
Software Stack and Technology Watch
Libraries
Module framework
Optimization techniques for scalability
Examples for application scalability description
Technology Watch
HP-SEE Bioinformatics portal
Deliverables
- D8.1 - Software scalability analysis and interoperability issues assessment
- D8.2 - Design of interoperability and scalability solutions
- D8.4 - Assessment of interoperability and scalability solutions
Nagios sensor for applications software and tools checking
https://hpseemon.ipb.ac.rs/nagios/cgi-bin/status.cgi?host=all
Dissemination and Training
HPC Initiatives, Procurement and Organization Guidelines
National HPC task-force modeling and organizational guidelines
Templates
- Project Logo
[PNG]